Leukocytes
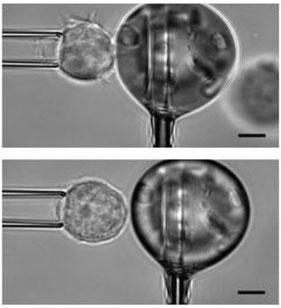
We have developped a micropipette force probe that allows us to measure piconewton to nanonewton-forces. We measure forces generated by T lymphocytes in contact with antibody-covered microbeads that mimick antigen presenting cells or targets for cytotoxic T cells. The technique allows us to dissect the mechanical response of leukocytes depending on the engaged receptors and on T cell type and/or modification. This tool also allows probing how leukocytes are sensitive to the mechanical properties of their environment.

Science Signaling, 2020; 13(627),
eaaw8214. Diacylglycerol kinase z regulates actin
cytoskeleton remodeling and mechanical forces at the B cell immune synapse. Sara V. Merino-Cortes, Sofia R.
Gardeta, Sara Roman-Garcia, Ana Martínez-Riaño, Judith Pineau, Rosa
Liebana, Isabel Merida, Ana-Maria Lennon Dumenil, Paolo Pierobon,
Julien Husson, Balbino Alarcon, and Yolanda R. Carrasco.
Cellular Microbiology, 2020; e13166. Shigella impairs human T lymphocyte responsiveness by hijacking actin cytoskeleton dynamics and TCR vesicular trafficking. Fatoumata Samassa, Mariana L. Ferrari, Julien Husson, Anastassia Mikhailova, Ziv Porat, Florence Sidaner, Katja Brunner, Teck-Hui Teo, Elisabetta Frigimelica, Jean-Yves Tinevez, Philippe J. Sansonetti, MariaIsabel Thoulouze, Armelle Phalipon.

Molecular Biology of the Cell, 2017; 28(23): 3229-3239. Micropipette Force Probe to quantify single-cell force generation: application to T cell activation. A. Sawicka, A. Babataheri, S. Dogniaux, A. I. Barakat, D. Gonzalez-Rodriguez, C. Hivroz, and J. Husson.
Cell, 2016;165(1):100-110. Cytotoxic T cells use mechanical force to potentiate target cell killing. Basu R*,
Whitlock BM*, Husson J*, Le Floc’h A, Jin W, Dotiwala F, Giannone G, Hivroz C, Lieberman J, Kam LC, and Huse M. (* co-first authors)
Molecular Biology of the Cell, 2016; 27(22): 3574-3582. Guillou L, Babataheri A, Saitakis M, Bohineust A, Dogniaux S, Hivroz C, Barakat AI, and Husson J. T lymphocyte passive deformation is controlled by unfolding of membrane surface reservoirs.
Force generation upon T cell receptor engagement.
Husson J, Chemin K, Bohineust A, Hivroz C, Henry N.
Langmuir, 2012;28(14):6106-13. Biomimetic droplets for artificial engagement of living cell surface receptors: the specific case of the T-cell. Bourouina N, Husson J, Hivroz C, Henry N.
Soft Matter, 2011;7:9130-9139. Formation of specific receptor–ligand bonds between liquid interfaces. Bourouina N, Husson J, Waharte F, Pansu RB, Henry N.
Endothelial Cells
Biophysical Journal, 2015; 109(2):209-19. Hogan B, Babataheri A, Hwang Y, Barakat AI, Husson J. Characterizing Cell Adhesion by Using Micropipette Aspiration.
Scientific Reports, 2016;6:21529. Dynamic monitoring of cell mechanical properties using profile microindentation. Guillou L, Babataheri A, Puech P-H, Barakat AI, and Husson J

Biophysical Journal, 2016; 111(12):2711-2721. D. Gonzalez-Rodriguez*, L. Guillou*, F. Cornat, J. Lafaurie-Janvore, A. Babataheri, E. de Langre, A. I. Barakat, and J. Husson. Mechanical criterion for the rupture of a cell membrane under compression.
Bacteria

Cell,
2018;174(1):143-155. Intermittent pili-mediated forces fluidize Neisseria meningitidis aggregates promoting vascular colonization. D. Bonazzi, V.
Lo Schiavo, S. Machata, I. Djafer-Cherif, P. Nivoit, V. Manriquez, H. Tanimoto, J. Husson, N. Henry, H. Chaté, R. Voituriez, and G. Duménil.
Mitochondria
Mitochondria constantly undergo fission and fusion events. These dynamical processes regulate mitochondrial morphology and are essential for cell physiology. We propose an elastocapillary mechanical instability as a mechanism for mitochondrial fission. We induce mitochondrial fission by rupturing the cell’s plasma membrane with a micropipette. We present a stability analysis that successfully explains the observed fission wavelength and the role of mitochondrial morphology in the occurrence of fission events.
PRL 2015;115:088102. Gonzalez-Rodriguez D, Sart S, Babataheri A, Tareste D, Barakat AI, Clanet C, Husson J. Elastocapillary Instability in Mitochondrial Fission
Previous work
Cell, 2012; 148(3):502-14. Laan L, Pavin N, Husson J, Romet-Lemonne G, van Duijn M, López MP, Vale RD, Jülicher F, Reck-Peterson SL, Dogterom M. Cortical dynein controls microtubule dynamics to generate pulling forces that position microtubule asters.
Biophysical Reviews and Letters, 2009; 4:33-44. Husson J, Laan L, Dogterom M. Force-generation by microtubule bundles.
PNAS, 2008; 105(26):8920-5. Laan L, Husson J, Munteanu EL, Kerssemakers JW, Dogterom M. Force-generation and dynamic instability of microtubule bundles.
Biophysical Journal, 2005; 89(6):4374-81. Pincet F, Husson J. The solution to the streptavidin-biotin paradox: the influence of history on the strength of single molecular bonds.
The Journal of chemical physics, 2009; 130(5):051103. Husson J, Dogterom M, Pincet F. Force spectroscopy of a single artificial biomolecule bond: the Kramers’ high-barrier limit holds close to the critical force.
Cellular and Molecular Bioengineering, 2008; 1(4):263-275. Gourier C, Jegou A, Husson J, Pincet F. A Nanospring Named Erythrocyte. The Biomembrane Force Probe.
PRE, 2008; 77(2 Pt 2):026108. Husson J, Pincet F. Analyzing single-bond experiments: influence of the shape of the energy landscape and universal law between the width, depth, and force spectrum of the bond.
